

GNN Species Maps
|
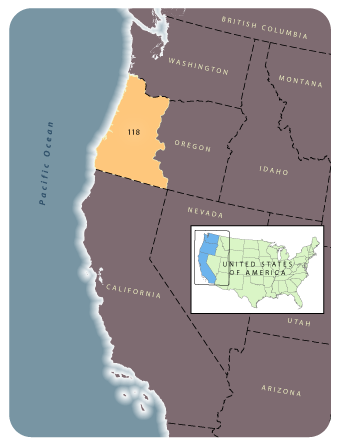
OverviewThis page provides links for downloading GNN species maps. For modeling region 118 (see map), a single GNN model was developed for the entire area (see Ohmann et al. 2011). Response variables used in model development were cover by species for woody species (all tree species, and shrub species present on at least 20 plots). This species model excludes satellite imagery, disturbance, and land ownership variables as explanatory variables, as they are more strongly correlated with forest structure than with species composition. |
Map Products
Digital GNN imputation maps are provided as 30-m-resolution ArcGIS grids, where the grid value is a unique plot number that links to the plot database. Selected vegetation variables from the plot database are joined as items in the grid to facilitate viewing and exploratory spatial analysis. Metadata for the vegetation variables are included with the grids and in the plot database.
The GNN models apply only to forest land (areas currently or with the potential to support at least 10% tree cover), because a consistent regional plot sample of nonforest areas is unavailable. Most users will want to use "masked" versions of the GNN maps (provided below), where areas of nonforest land cover developed from the GAP Analysis Program's Ecological Systems map have been embedded in the GNN grids. Unmasked versions of the GNN maps are available upon request, for users who would like to apply a nonforest mask of their own choosing.
The accuracy assessment reports included in the download file are compiled at the modeling region level and provide accuracy assessment information for plots within each modeling region.
| Product Description | Status | Imagery year | Date posted | Download |
|---|---|---|---|---|
| GNN species map and accuracy assessment report | FINAL | N/A | 02/23/2011 |
Data Dictionary
The data dictionary below lists all fields that have been attached to the above map products. If a field has coded entries, clicking on the plus symbol in the first column ('C') will show all codes. For information on how a field is calculated, click on the plus symbol in the second column ('F') for details. The dictionary can be fully expanded or collapsed with the below links.